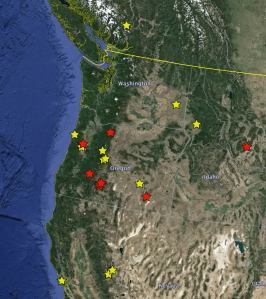
In September of 2009, I arrived in Oregon, excited to begin my new position at Oregon State University. I was also excited to live near Marys Peak, as the top of Marys Peak was the locality of capture of the only known specimen of what was then called Bembidion chintimini. (I’ve since moved that and related species out of the genus Bembidion, and as the genus Lionepha.) A few days after I arrived in Corvallis, I drove up to the top of that mountain to see if I could find the species again. I found some little beetles that might or might not be Bembidion chintimini, but I was a novice with that group of bembidiines, and so I wasn’t sure. I also found a larger, related specimen lower down the slopes of Marys Peak, near Alder Creek Falls, which confused me as well. My efforts to discover the identity of those specimens took a long time, and the threads I was tugging on as I explored caused an unravelling in my understanding of that group of beetles, which I would eventually re-weave into a new tapestry. In the end, this path led to the discovery of a total of four new species, as well as a merging of three others.

In many ways this project was the heart and soul of my first decade in Oregon. I spent many hundreds of hours on this project, traveling and collecting specimens, extracting and sequencing their DNA, making genitalic preparations, studying their structures, photographing them, examining their chromosomes, doing phylogenetic analyses, and, eventually, creating the figures for the paper and writing the text. Perhaps more emotionally compelling was the discovery and solving of many puzzles along the way, especially that revolving around “Bembidion chintimini”. That some of these puzzles were solved by a combination of old-fashioned morphological studies as well as next-generation genomic sequencing of old type specimens, including a 159-year-old LeConte lectotype, made the tale all the more compelling to me.
John Sproul, a former graduate student of mine, helped by sequencing the DNA I extracted from some pinned type specimens (including that LeConte lectotype), as well as by doing important collecting in the Sierra Nevadas of California; for these efforts, he is a co-author of the paper that has finally resulted.
This paper, titled “Species delimitation, classical taxonomy and genome skimming: a review of the ground beetle genus Lionepha (Coleoptera: Carabidae)“, came out recently in the Zoological Journal of the Linnean Society. The paper can be found at https://doi.org/10.1093/zoolinnean/zlz167; if you would like a PDF, email me.
It turns out that I did find a female of “Bembidion chintimini” on Marys Peak that first trip in 2009. It took until the following summer for me to realize that. It took even longer to eventually come to the realization that this species was widespread, and had a much older name (Lionepha erasa LeConte). Here’s a condensed version of the story, as told in the paper:
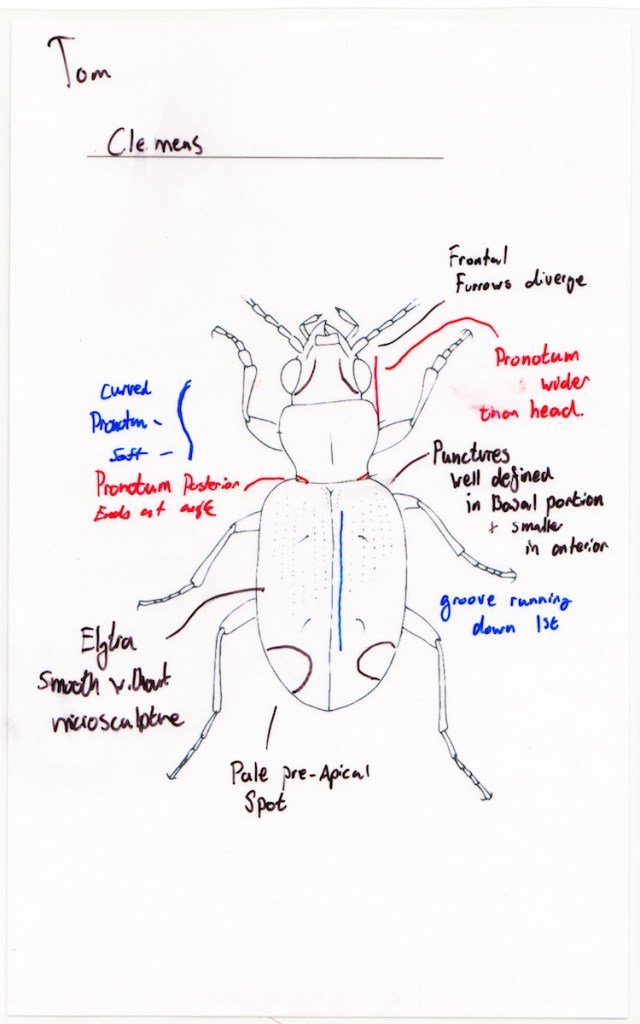
Investigation of the rarer species, the one here called Lionepha erasa, began in 2010. Dissection of the first recognized males from Marys Peak, Oregon (type locality of Bembidion chintimini) revealed an aedeagus indistinguishable from those from San Juan Island, Washington (type locality of Bembidion lummi). The female holotype of B. chintimini is wingless and has slightly rounded shoulders. However, the Marys Peak population is wing-dimorphic, and winged individuals are in body form no different from the type series of Bembidion lummi. The elytral microsculpture of the holotype of B. chintimini is perfectly isodiametric (against Erwin & Kavanaugh, 1981), thus matching that of B. lummi. Other characters mentioned by Erwin & Kavanaugh as distinguishing the two populations are not consistent with available specimens. The lack of evident morphological differences, combined with effectively identical DNA sequences in specimens from Oregon, British Columbia and Alaska suggested that the Marys Peak populations are the same species as populations further north, and for this reason, Bembidion chintimini and B. lummi were synonymized by Maddison in Kanda et al. (2015).
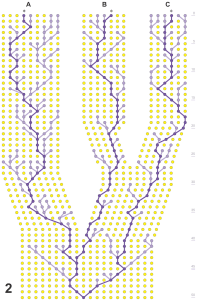
This left in question the specimens considered to be Bembidion lindrothellus by Erwin & Kavanaugh, which are at first glance similar to the Marys Peak and other populations of ‘Bembidion chintimini’. Specimens classified as Bembidion lindrothellus are reported to be paler, but all specimens mentioned in Erwin & Kavanaugh (1981) are teneral. The unsclerotized aedeagus of the holotype of Bembidion lindrothellus made comparison of internal sac sclerites difficult. However, the internal sac membrane that rests in the left-most position has a species-specific microsculpture in Lionepha, and the microsculpture scales of the holotype of Bembidion lindrothellus from Alaska match those of Marys Peak specimens. A non-teneral male was also collected by Lindroth at the type locality of B. lindrothellus, but was not included in the type series, perhaps as the specimen was housed in Lindroth’s collection in Lund, Sweden. This specimen is presumably the one whose genitalia Lindroth figured as Bembidion brumale (1963: fig. 127f). We have examined that specimen, and it is indistinguishable from specimens of ‘Bembidion chintimini’ from Alaska, British Columbia, Washington and Oregon, including details of the internal sac. Most critically, DNA sequences of the holotype of Bembidion lindrothellus are identical in eight studied genes to those of other specimens from throughout the range (Figs 5–7). It is thus evident that the holotypes of Bembidion chintimini, B. lindrothellus and B. lummi belong to a single species.Maddison and Sproul (2020)
However, there is an older name. The type series of Bembidium erasum consists of four females. These specimens have traditionally been considered to belong to the common, widespread species here called Lionepha probata. Females of these two isodiametrically microsculptured species are difficult to tell apart, especially those with less-extreme prothoracic proportions (neither wide nor narrow). Although there are distinctions in the lobe of the female bursa of fully sclerotized individuals, interpretation of tenerals is more tenuous. Specimens in the type series of Bembidium erasum are all teneral, with prothoraces of moderate width, and thus there is no clear morphological evidence to place them to species. The type series was provided by George Suckley (LeConte, 1859), presumably captured during his travels as naturalist for the governor of Washington Territory during 1853–57 (Cooper & Suckley, 1859). The type series is from ‘Oregon’, which at the time encompassed the current area of Oregon, the southern half of what is now Idaho and some parts of Wyoming and Montana (Barry, 1932). Suckley’s travels in Oregon included areas within the range of both species (Cooper & Suckley, 1859), and thus geography provides no clues about species membership. However, DNA data from the lectotype (and two of the paralectotypes; Sproul & Maddison, 2017) makes it clear that these specimens belong to the current species (Figs 5–7; Supporting Information, Fig. S1). Thus, the valid name of this species is Lionepha erasa, with Bembidion chintimini, B. lindrothellus and B. lummi as junior synonyms.
That description of the history does not adequately capture all the many mysteries, proposed and rejected explanations, and confirmed hypotheses along my multi-year path, and the eventual pleasure as the hypotheses became confirmed through the emergence of consistent, repeated patterns. It was one of the more pleasing journeys of discovery I have been on. I’ve previously mentioned a few of the turns and twists in the story, in my posts on Rainy-season beetles and surprises in Lionepha.
In addition to solving the mystery of the small beetles on top of Marys Peak, four new species were discovered along the way and described in the paper:
- The larger specimen I discovered on my first trip up Marys Peak belongs to a species that lives along the sides of Alder Creek Falls and on rock seeps in the area, as well as along a creek west of Eugene, Oregon, and in the Trinity Alps of California, is now officially named Lionepha tuulukwa Maddison. I’ve talked about the naming of this beetle here and here.
- A species I first became aware of through a single male along Bishop Creek that John Sproul found, is now called Lionepha lindrothi Maddison & Sproul.
- The species I found in the central Sierra Nevada of California, and which provided another surprise in the group, is now called Lionepha australerasa Maddison.
- A species from western Montana and Wyoming, as well as eastern Oregon and Washington, came to light from specimens collected by my good friend David Kavanaugh. This species is now called Lionepha kavanaughi Maddison.
This paper on Lionepha implicitly tells a love story between me and my beetles. When I look at the final product, I see it as a celebration of the process and fruits of discovery. I also see it as a history of my embracing the Pacific Northwest, and the abundant and diverse life that lives here.